 Analysis of MiSeq Dataįor bioinformatic analysis of MiSeq data, we suggest using Purple line in DNA Subway ( ). fa, and you will need to change the command accordingly. The extension for the sequence file might be. Then, type “cat *.fasta > combined.fasta”. To combine fasta files on a Mac, navigate to the folder with the files in the Terminal. 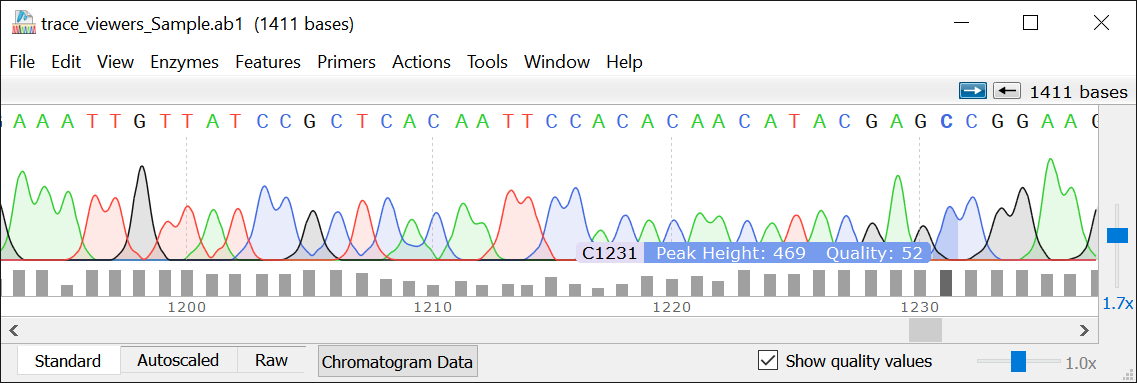
For directions on how to combine fasta files (which are text files) in Windows, see. The sequences for each individual colony can be submitted separately or the fasta files for each colony can be combined into a single fasta file first. More accurate identification is possible using a curated database of 16s rRNA sequences, such as Silva ( ). For more details, see our Student handout on BLAST analysis of sequencing data. Sequences can be trimmed in SnapGene Viewer and then exported as fasta files.īacterial species (or more typically genera) can be identified using a BLAST search on Genbank. 
An alternative is to have students view the chromatograms and quality data in a viewer such as SnapGene Viewer ( ). Sanger sequencing of colony-based PCR products will result in chromatogram files and sequence (or fasta) files.Ĭhromatogram files in standard PDF format can be viewed and fasta files trimmed in a text editor. 
Analysis of colony-based PCR 16s rRNA data 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
